Note
Click here to download the full example code
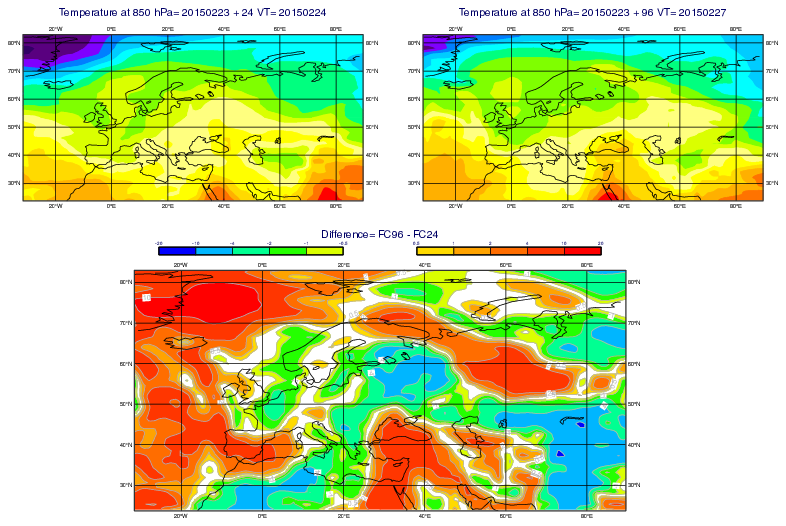
GRIB - Layout with 3 Maps
# (C) Copyright 2017- ECMWF.
#
# This software is licensed under the terms of the Apache Licence Version 2.0
# which can be obtained at http://www.apache.org/licenses/LICENSE-2.0.
#
# In applying this licence, ECMWF does not waive the privileges and immunities
# granted to it by virtue of its status as an intergovernmental organisation
# nor does it submit to any jurisdiction.
#
# ------------------------------------------------------------------
# Demonstrates how to use a Display Window icon to
# produce a plot with 3 maps consisting of a) a 24h
# forecast, b) a 96h forecast and c) their difference
# ------------------------------------------------------------------
import metview as mv
# read the 24h forecast GRIB data from file
filename = "t_fc24.grib"
if mv.exist(filename):
f24 = mv.read(filename)
else:
f24 = mv.gallery.load_dataset(filename)
# read the 96h forecast GRIB data from file
filename = "t_fc96.grib"
if mv.exist(filename):
f96 = mv.read(filename)
else:
f96 = mv.gallery.load_dataset(filename)
# filter just the 850hPa level
t_fc24 = mv.read(data=f24, levelist=850)
t_fc96 = mv.read(data=f96, levelist=850)
# we will use the same view in all the plots, but we could also use different
# views for each plot
view = mv.geoview(
map_area_definition="corners", area=[23.7, -31.6, 83.0, 89.6] # rough European area
)
# each page in the layout contains its position plus its view specification
# (could also be a Cross Section View, for example). Defaults are=
# top=0, bottom=100, left=0, right=100 in percent of layout height and width
page_24 = mv.plot_page(bottom=45, right=50, view=view)
page_96 = mv.plot_page(bottom=45, left=50, view=view)
page_diff = mv.plot_page(top=50, left=11, view=view)
# contouring specifications
t_shade = mv.mcont(
contour="off",
contour_hilo="off",
contour_label="off",
contour_level_selection_type="interval",
contour_interval=4,
contour_shade="on",
contour_shade_min_level=-52,
contour_shade_max_level=48,
contour_shade_method="area_fill",
contour_shade_colour_method="list",
contour_shade_colour_list=[
"rgb(0.88,0.2,0.88)",
"rgb(0.68,0.2,0.68)",
"rgb(0.48,0.2,0.48)",
"rgb(0.28,0.2,0.28)",
"rgb(0.2,0,0.4)",
"rgb(0.35,0,0.5)",
"blue_purple",
"greenish_blue",
"rgb(0,0.8,1.0)",
"blue_green",
"bluish_green",
"yellow_green",
"greenish_yellow",
"rgb(1,1,0.5)",
"yellow",
"orange_yellow",
"yellowish_orange",
"rgb(1,0.45,0)",
"red",
"rgb(0.8,0,0)",
"burgundy",
"rose",
"magenta",
"rgb(1,0.5,1)",
"rgb(1,0.75,1)",
],
)
pos_shade = mv.mcont(
legend="on",
contour_line_colour="grey",
contour_highlight="off",
contour_level_selection_type="level_list",
contour_level_list=[0.5, 1, 2, 4, 10, 20],
contour_shade="on",
contour_shade_method="area_fill",
contour_shade_max_level_colour="red",
contour_shade_min_level_colour="orange_yellow",
contour_shade_colour_direction="clockwise",
)
neg_shade = mv.mcont(
legend="on",
contour_line_colour="grey",
contour_highlight="off",
contour_level_selection_type="level_list",
contour_level_list=[-20, -10, -4, -2, -1, -0.5],
contour_shade="on",
contour_shade_method="area_fill",
contour_shade_max_level_colour="greenish_yellow",
contour_shade_min_level_colour="blue",
contour_shade_colour_direction="clockwise",
)
# when we have multiple pages in a layout, the default titles can be a bit too long
# for the available space; hence we will construct shorter titles, using automated
# fields as far as possible. We could also use Metview's own date/string formatting
# routines to construct 'nicer' dates in the titles
title_fc = mv.mtext(
text_line_1="<grib_info key='name'/> at <grib_info key='level'/> hPa= "
+ "<grib_info key='dataDate'/> + <grib_info key='step'/>"
+ " VT= <grib_info key='validityDate'/>",
text_font_size=0.45,
)
title_diff = mv.mtext(text_line_1="Difference= FC96 - FC24", text_font_size=0.45)
dw = mv.plot_superpage(
# the order of these pages is used when indexing them in the plot() command
pages=[page_24, page_96, page_diff]
)
# define the output plot file
mv.setoutput(mv.pdf_output(output_name="layoutx3"))
# plot the data into each page using a single plot command; note that
# we defined 3 pages, so they are indexed by 0, 1, 2
mv.plot(
dw[0],
t_fc24,
t_shade,
title_fc,
dw[1],
t_fc96,
t_shade,
title_fc,
dw[2],
t_fc96 - t_fc24,
neg_shade,
pos_shade,
title_diff,
)